One of the biggest challenges in modern medicine is the appearance of antibiotic-resistant bacteria. To combat this threat, there is an urgent need for novel antibiotics. Many antibiotics used today are only modified versions of existing drugs to which resistance quickly develops. Dr Scott Champney at East Tennessee State University has investigated a new bacterial antibiotic target, ribosome biosynthesis, that can be used to guide the development of novel antibiotics.
Antibiotic resistance poses a major public health threat. It occurs when bacteria evolve strategies to evade drugs which would previously have been fatal to them. Bacterial infections such as MRSA (methicillin-resistant Staph aureus) and multidrug-resistant tuberculosis are therefore becoming harder to treat. Antibiotic resistance manifests itself in increases in the length of time people stay in hospital, the cost of their treatment and in overall mortality.
There are several strategies to combat this challenge. The first involves changes around behaviour to help stop the development of infection in the first place, such as good hand and food hygiene and vaccination. There is also a push for smarter prescribing, where specific antibiotics should be used to treat a bacterium that is susceptible to them since oversubscription of broad-spectrum or non-effective antibiotics encourages the development of resistant bacteria. Finally, novel antibiotics are required, along with additional unexplored targets.
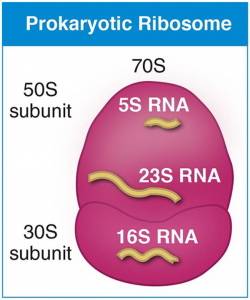
A major target for many antimicrobials is the bacterial ribosome. The 70S ribosome is a complex structure which is the catalytic site for protein synthesis. It has two subunits, the 30S with 16S ribosomal RNA and 21 ribosomal proteins and the 50S subunit with 23S and 5S RNAs and 35 proteins (see figure 1). Antibiotics like azithromycin bind to the 50S subunit and stop peptide bond formation between two amino acids. Compounds like neomycin attach to the 30S subunit and prevent the initiation of the protein synthesis process (see figure 2).
The research of Dr Scott Champney at East Tennessee State University has identified a new target for antibiotics which prevent the subunit assembly process. Subunit assembly is inhibited if an antibiotic binds to RNA in the precursors required for the formation of mature ribosomal subunits.
“A major target for many antibiotics is the bacterial ribosome.”
Subunit assembly
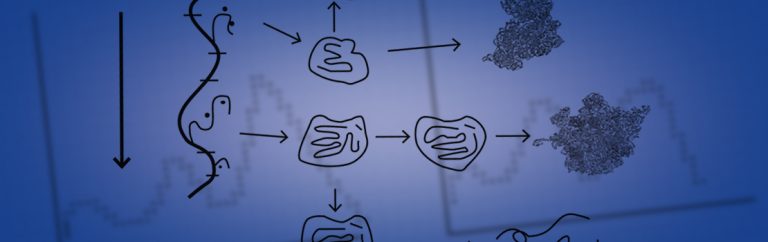
The bacterial ribosome is the structure most frequently targeted by current antibiotics. Ribosomal subunit biogenesis proceeds through an intermediate structure for each subunit (see figure 3). A 21S precursor for the 30S subunit is a check-point for proper assembly. This particle contains about one-half of the appropriate subunit proteins and a precursor 16S RNA which requires a major conformational change in order to mature into the functional structure. 50S subunit formation proceeds though 32S and 43S intermediates which also have a subset of the total proteins and also involves significant 23S RNA structural rearrangements. Chaperone proteins facilitate this process and ribonuclease enzymes trim the 5’ and 3’ ends of the three RNAs.

Dr Champney’s research has identified a novel target for many antibiotics, the inhibition of subunit assembly by drug binding to the precursor particle for each subunit (see figure 3). Ribosomal antimicrobials thus bind to RNA in the mature subunit to stop translation and to the same RNA site in the intermediate structure to prevent subunit assembly. Stalled assembly intermediates are then degraded by specific ribonucleases.
Experimental assays
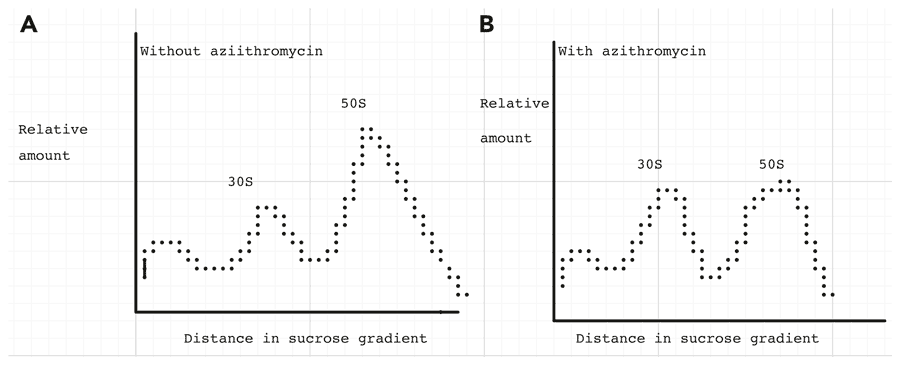
Dr Champney and his team utilised two different experimental assays to evaluate this novel target. Firstly, they used a unique four-part assay to characterise the antibiotic concentrations causing inhibition of the formation of the affected subunit. This involved a process called sucrose density gradient ultracentrifugation, which allows a mixture of different-sized molecules to be sorted in order from highest to lowest density (see figure 4). This technique allowed the researchers to quantify the amounts of each subunit structure present, for example the precursor subunits versus the mature subunits, as well as assessing the binding distribution of radioactively labelled antibiotics. The four-part assay also measured the concentration-dependent inhibition of the cellular growth rate, viability, protein synthesis rate and reduction in the amount of the affected subunit.
The researchers also used a pulse and chase radio-isotope labelling procedure to measure the effect of a particular antibiotic on the rate of subunit formation. This involves sampling with time the rate of formation of each subunit by using a radio-active RNA precursor molecule. The reduction in time for subunit formation was then assayed by the sucrose density gradient ultracentrifugation procedure.

They were able to show that 30S subunit formation was prevented by 3 aminoglycoside antibiotics, neomycin, paromomycin and hygromycin B. 50S subunit formation was inhibited by 45 unique antibiotics including macrolides, ketolides, lincosamides and streptogramins. The different pathogenic microorganisms examined included E.coli, Haemophilus influenzae, MRSA and drug-resistant Streptococci.
“The model proposed by Dr Champney predicts that antibiotics bind to ribosomal RNA in intermediate structures formed during the subunit assembly process.”
The investigations also helped to explain the slow recovery of inhibited cells after drug removal by showing that re-synthesis of the damaged subunit limited the recovery period. Also, the 50S subunit precursor was shown to be the natural substrate for an RNA methylating enzyme which confers antibiotic resistance to 50S-binding drugs. Finally, it was demonstrated that RNA-processing ribonucleases were novel inhibitory targets for a unique molecule, the vanadyl ribonucleoside complex. Significantly, this inhibitor blocked the assembly of both subunits but did not impair translation. A search for other ribonuclease inhibitors should be fruitful.
A hypothetical model
While further research is needed to fully understand the way by which these drugs work, Dr Champney proposes a hypothetical model by which most of the antibiotics tested can inhibit subunit assembly and translation. The model proposed by Dr Champney predicts that antibiotics bind to the ribosomal RNA in intermediate structures formed during the assembly process (see figure 3). This subsequently prevents subunit maturation. Enzymes then degrade the RNA in the unsuccessful precursor molecule. He suggests that there are a number of different ways antibiotics can inhibit the subunit assembly process, dependent on where they bind.
Dr Champney has identified a novel antibiotic target that can be further examined in the hunt for new antibiotics and other novel targets. The identification of more ribonuclease inhibitors may present opportunities for the development of strategies to combat antibiotic resistance.
Is there a danger that bacteria will start to evolve ways to evade ribosomal antibiotics in the future?
This is a distinct possibility since selection for drug-resistant microorganisms is constently occurring. This is why the development of unique antibiotics and the identification of new targets is essential.
References
- Champney, W.S. (2020). Antibiotics targeting bacterial ribosomal subunit biogenesis. The Journal of antimicrobial chemotherapy, 75(4), 787–806. Available at: https://doi.org/10.1093/jac/dkz544
- Frazier, A.D. and Champney, W.S. (2012). The vanadyl ribonucleoside complex inhibits ribosomal subunit formation in Staphylococcus aureus. The Journal of antimicrobial chemotherapy, 67(9), 2152–2157. Available at: https://doi.org/10.1093/jac/dks182
- Foster, C. and Champney, W.S. (2008). Characterization of a 30S ribosomal subunit assembly intermediate found in Escherichia coli cells growing with neomycin or paromomycin. Archives of microbiology, 189(5), 441–449. Available at: https://doi.org/10.1007/s00203-007-0334-6
- Pokkunuri, I. and Champney, W.S. (2007). Characteristics of a 50S ribosomal subunit precursor particle as a substrate for ermE methyltransferase activity and erythromycin binding in Staphylococcus aureus. RNA biology, 4(3), 147–153. Available at: https://doi.org/10.4161/rna.4.3.5346
- Champney, W.S. and Tober, C.L. (1999). Molecular investigation of the postantibiotic effects of clarithromycin and erythromycin on Staphylococcus aureus cells. Antimicrobial agents and chemotherapy, 43(6), 1324–1328. Available at: https://doi.org/10.1128/AAC.43.6.1324
10.26904/RF-133-3033
Research Objectives
Dr Champney has identified and characterised a novel target for ribosomal antibiotics.
Funding
This research was funded by the National Institutes of Health and by seven pharmaceutical companies.
Collaborators
- Dr Justin Beach
- Dr Harold Chittum
- Dr Cerrone Foster
- Dr Ashley Frazier
- Dr Susan MaGaha
- Roopal Mehta
- Dr Indira Pokkunuri
- Jessica Silvers
- Craig Tober
- Dr Jerry Usary
Bio
Dr Scott Champney is an Emeritus Professor in the Department of Biomedical Sciences in the Quillen College of Medicine at East Tennessee State University. He has a PhD from SUNY Buffalo and has been on the faculty of the University of Georgia, the University of Texas at San Antonio and at East Tennessee State University for the past 31 years. He has taught medical students and PhD graduate students and has conducted a grant-funded research program. His research interests include investigating the inhibitory effects of antibiotics on bacterial ribosome formation. He has 75 research publications and a book, New Antibiotic Targets (Humana Press, 2008).


Contact
Dr Scott Champney
E: Champney@etsu.edu
T: +1 423 202 6977