There are roughly as many bacterial cells in the human body as there are human cells, with approximately 1,014 bacteria in an average human large intestine alone. In recent years, there has been increasing interest in the relationship between the bugs that live in the human intestine, the gut microbiome, and human health and disease. Dr Casey Theriot’s work mainly focuses on one enteric pathogen, Clostridium difficile, and its relationship with the resident gut microbiota. Dr Theriot focused on this pathogen because it is exquisitely sensitive to changes in the gut microbiota and metabolome, meaning it is a good bacterium to look at in experimental settings.

The battle of the bugs
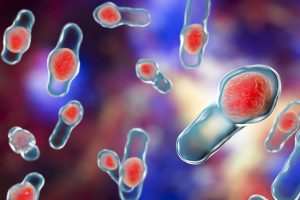
C. difficile is an anaerobic, spore-forming, gram-positive bacteria that was first discovered in 1935. C. difficile spores require specific bile acids for maximal germination into metabolically active vegetative cells. Once reaching this state, the C. difficile cells require amino acids and carbohydrates to grow to high cell density, produce toxins, which mediates disease.
Knowledge of the mechanisms by which the gut microbiota confers colonisation resistance against C. difficile is incomplete, meaning that it is difficult to improve preventative and therapeutic approaches against this pathogen. Dr Theriot is currently working on a project, which aims to define members of the gut microbiota that are able to influence C. difficile colonisation and pathogenesis.
Dr Theriot and her group want to address two questions. Firstly, whether restoring microbial-mediated secondary bile acid metabolism in the large intestine restores colonisation resistance against C. difficile and secondly, whether restoring bacteria that are able to compete with C. difficile for nutrients required for growth, can re-establish colonisation resistance against C. difficile infection.
There are approximately as many bacterial cells in the human body as there are human cells, with approximately 1014 bacteria in an average human large intestine alone.
To answer the first question, the team will characterise bacteria that are capable of secondary bile acid metabolism. Genetic engineering of bacterial strains for efficient enzyme delivery to the gastrointestinal tract will be evaluated in an experimental laboratory setting and in a mouse model of CDI. The team will use both bacterial expression systems and the revolutionary CRISPR-Cas9 gene editing system to achieve this, allowing an optimum and controllable delivery system that would allow secondary bile acids to be produced.
It is not a new idea that members of the gut bacterial community compete with each other for nutrients, as well as competing with pathogenic bacteria; there is competition between the resident gut microbiota too.
Previously in 1988, it was first highlighted how the colonic microbiota was able to compete for nutrients, including glucose and sialic acids, with C. difficile, resulting in suppression of growth of C. difficile. If it is possible to identify the groups of bacteria, which compete with C. difficile, there is the potential to use these as a therapeutic for competitive exclusion of pathogens by the resident gut microbiota. Therefore, to address the second question, bacteria that are able to compete for the same nutrients as C. difficile will be characterised. These bacteria will then be evaluated using competition assays in the laboratory and in a mouse model of CDI.
All about bile acids
Bile acids are synthesised from cholesterol by liver enzymes. Once synthesised, the primary bile acids travel through the small intestine, where 95% of bile is absorbed. The small amount of bile acids that reach the large intestine are further converted into secondary bile acids by members of the gut microbiota.
Work from Dr Theriot has shown that providing there are no antibiotics present, many microbial-derived secondary bile acids are able to inhibit C. difficile growth. The group also showed that following antibiotic administration, secondary bile acid metabolism is depleted, permitting C. difficile growth, toxin production, and disease. In this study, mice were treated with various antibiotics, including metronidazole, vancomycin, and kanamycin. Some groups were allowed to recover from antibiotic treatment for one, two or six weeks, to investigate the time taken for resistance to C. difficile to re-establish in the gut and changes were examined in the small intestine and in the large intestine of the mice. In particular, they found that large intestinal content that was resistant to C. difficile had higher levels of secondary bile acids, and lower levels of primary bile acids, compared to large intestinal content that allowed for C. difficile germination and outgrowth. In particular, Dr Theriot and collaborators have shown that the bacterial families Lachnospiraceae and Ruminococcaceae in the large intestinal microbiota are positively correlated with secondary bile acids.
The production and consumption of certain metabolites (products of metabolic reactions), including secondary bile acids, amino acids and sugars, by the resident gut microbiota contribute to resistance against C. difficile. The rationale behind this is that certain bacteria produce products resulting in resistance to CDI and that removing these bacteria, and their metabolites, from the intestinal ecosystem, results in proliferation of pathogens such as C. difficile.
Targeted approaches provide the opportunity to use targeted engineering and editing of the microbiome to alter function, and consequently to prevent and treat human diseases.
Future directions
One current approach to restore the gut microbiota after antibiotic administration is the use of faecal transplantation. This process aims to repopulate the gut with healthy bacteria from a donor stool sample. Even though treatments like these work in 90% of cases, the mechanisms and long-term consequences of these treatments are still unclear, further emphasising the need for work, such as that of Dr Theriot, to establish targeted bacterial therapies to restore colonisation resistance against C. difficile. Dr Theriot’s work will contribute to understanding how the gut microbiota regulates bile acid, amino acid, and carbohydrate metabolism in the large intestine, another aspect that is also crucial in the fight against CDI. The approach taken by Dr Theriot’s group is innovative because it focuses on the use of targeted bacterial therapy to restore secondary bile acids and bacterial competition, ultimately leading to the restoration of colonisation resistance against C. difficile in the gastrointestinal tract. It would provide alternative therapeutic approaches, leading to a move away from antibiotic use, which is also in itself a risk factor for CDI. Furthermore, C. difficile represents just one specific pathogen; once the system is established then targeted approaches could be used against many other pathogens in the future, providing the opportunity to use targeted engineering and editing of the microbiome to alter function, and consequently to prevent and treat human diseases.
The information gained by the proposed research is expected to lead to novel approaches for the prevention and treatment of C. difficile infection. The next step in evaluating candidate bacterial therapeutics from this proposal is translational studies in humans. I am currently building partnerships with clinicians at UNC and Duke Hospital in Research Triangle to work toward reaching this goal in the near future. It is my hope that this targeted probiotic therapeutic approach will be used for modulation and treatment of many gastrointestinal diseases, and metabolic diseases like obesity and diabetes.
References
- https://theriotlab.org/
- https://cvm.ncsu.edu/directory/theriot-casey/
- http://grantome.com/grant/NIH/R35-GM119438-01/
- Theriot, CM. 2018. Beyond structure: defining the function of the gut using omic approaches for rational design of personalized therapeutics. mSystems 3:e00173 17. doi: 10.1128/mSystems.00173-17.
- Theriot, CM, et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nature Communications. 5:3114 doi:10.1038/ncomms4114 (2014).
Dr Theriot’s current research focuses on manipulating the gut microbiota to rationally alter the composition of the bile acid pool in the gut, and restore microbial competition, which has the potential to improve preventative and therapeutic approaches against many human diseases.
Funding
- NIH National Institute for General Medicine NIGMS
- NCSU CVM NC State University College of Veterinary Medicine, Raleigh, NC
- UNC CGIBD Center for Gastrointestinal Biology and Disease, University of North Carolina, Chapel Hill, NC
- NIH National Institute Diabetes and Digestive Kidney Diseases NIDDK
- UNC North Carolina Translational and Clinical Sciences Institute
Collaborators
- Vincent Young, MD, PhD, University of Michigan
- Anna Seekatz, PhD, University of Michigan
- Rodolphe Barrangou, MS, MBA, PhD, NC State University
- Sarah O’Flaherty, PhD, NC State University
- Stephanie Montgomery, DVM, PhD, University of North Carolina
- Ajay Gulati, MD, University of North Carolina
- Michael Dougherty, MD, University of North Carolina
- Sarah McGill, MD, University of North Carolina
- Cristina Lanzas, DVM, PhD, NC State University CVM
- Andrew Patterson, PhD, Penn State University
- The Theriot lab on the website : https://theriotlab.org
Bio
 Casey Theriot is an Assistant Professor in Infectious Diseases at NC State University College of Veterinary Medicine in Raleigh, NC. Dr Theriot received her PhD from NC State University in the Department of Microbiology. She then went on to complete a postdoctoral fellowship and independent research position with Dr Vincent Young at the University of Michigan Medical School.
Casey Theriot is an Assistant Professor in Infectious Diseases at NC State University College of Veterinary Medicine in Raleigh, NC. Dr Theriot received her PhD from NC State University in the Department of Microbiology. She then went on to complete a postdoctoral fellowship and independent research position with Dr Vincent Young at the University of Michigan Medical School.
Contact
Dr Casey M. Theriot, PhD
Assistant Professor Infectious Disease
College of Veterinary Medicine
Department of Population Health and Pathobiology
North Carolina State University
1060 William Moore Drive RB 406
Raleigh, NC 27607
USA
E: cmtherio@ncsu.edu
T: +1 919 513 0711
twitter: @TheRiotMicrobe
W: https://theriotlab.org/
W: https://cvm.ncsu.edu/directory/theriot-casey/









